What Is The Promoter, Terminator, Template Strand, Non-template Strand
2.ane: Overview of Transcription
- Page ID
- 32270
Learning Objectives
- Place the key steps of transcription, the part of the promoter and the function of RNA polymerase.
- Distinguish between coding (RNA-similar) and non-coding (template) strands of Deoxyribonucleic acid. Understand that within a single piece of DNA, either strand can be used as the template for dissimilar genes, merely the RNA volition still be produced from 5' → 3'.
- Draw a line diagram showing a segment of DNA from a gene and its RNA transcript, indicating which DNA strand is the template, the management of transcription and the polarities of all Deoxyribonucleic acid and RNA strands.
- Give examples of non-coding RNA molecules.
What is transcription?
Consider that all of the cells in a multicellular organism have arisen by division from a unmarried fertilized egg and therefore, all accept the same DNA. Sectionalisation of that original fertilized egg produces, in the case of humans, over a trillion cells, past the time a baby is produced from that egg (that's a lot of Deoxyribonucleic acid replication!). Even so, we also know that a baby is not a giant brawl of a trillion identical cells, but has the many dissimilar kinds of cells that brand up tissues like pare and muscle and bone and nerves. How did cells that take identical Dna turn out so unlike?
The answer lies in cistron expression, which is the process by which the data in DNA is used. Although all the cells in a babe have the same Dna, each different jail cell type uses a different subset of the genes in that Deoxyribonucleic acid to directly the synthesis of a distinctive set of RNAs and proteins. The first step in cistron expression is transcription, the process of copying information from Dna sequences into RNA sequences. This process is besides known as Dna-dependent RNA synthesis. When a sequence of Deoxyribonucleic acid is transcribed, only ane of the two DNA strands is copied into RNA, when this RNA encodes a poly peptide is it known as messenger RNA (mRNA).

Important features of transcription
- All RNA, mRNA as well as tRNA, rRNA, microRNA and more, is produced past transcription.
- Only ane strand of DNA is used as a template past enzymes called RNA polymerases
- RNA is synthesized from five' to 3'.
- RNA polymerases do not need primers to begin transcription.
- The iv ribonucleotide triphosphates (rNTPs) are ATP, GTP, UTP, and CTP.
- RNA polymerases brainstorm transcription at DNA sequences chosen promoters.
- RNA polymerases end transcription at sequences called terminators.
In transcription, an RNA polymerase uses only 1 strand of DNA, called the template strand, of a gene to catalyze synthesis of a complementary, antiparallel RNA strand. RNA polymerases employ ribose nucleotide triphosphate (NTP) precursors, in dissimilarity to Deoxyribonucleic acid polymerases, which utilise deoxyribose nucleotide (dNTP) precursors (compared on page ane.1: The Structure of DNA). In add-on, RNAs contain uracil (U) nucleotides into RNA strands instead of the thymine (T) nucleotides used in DNA. RNA polymerases differ from Deoxyribonucleic acid polymerases in that they do not require primers. With the help of transcription initiation factors, RNA polymerase locates the transcription offset site of a gene and begins synthesis of a new RNA strand from scratch by joining the ii ribonucleotides that are complementary to the starting time two bases of the template strand.
Overview of the Stages of Transcription
The basic steps of transcription are initiation, elongation, and termination. Here we tin place several of the DNA sequences that narrate a gene. The promoter is the binding site for RNA polymerase. Information technology unremarkably lies 5' to, or upstream of the transcription outset site. Binding of the RNA polymerase positions the enzyme to well-nigh the transcription start site, where it will start unwinding the double helix and brainstorm synthesizing new RNA. The transcribed gray DNA region in each of the three panels are the transcription unit of the gene. Termination sites are typically 3' to, or downstream from the transcribed region of the gene. By convention, upstream refers to Deoxyribonucleic acid 5' to a given reference bespeak on the Deoxyribonucleic acid (e.thou., the transcription start-site of a gene). Downstream and then, refers to Deoxyribonucleic acid 3' to a given reference betoken on the Deoxyribonucleic acid.
RNA polymerase
Edifice an RNA strand is very like to edifice a DNA strand. This is not surprising, knowing that DNA and RNA are very like molecules. What enzyme carries out transcription? Transcription is catalyzed by the enzyme RNA Polymerase. "RNA polymerase" is a general term for an enzyme that makes RNA. There are many different RNA polymerases.

Like Dna polymerases, RNA polymerases synthesize new strands just in the 5' to 3' management, but because they are making RNA, they use ribonucleotides (i.due east., RNA nucleotides) rather than deoxyribonucleotides. Ribonucleotides are joined in exactly the same way as deoxyribonucleotides, which is to say that the 3'OH of the terminal nucleotide on the growing chain is joined to the 5' phosphate on the incoming nucleotide.
1 important departure between Deoxyribonucleic acid polymerases and RNA polymerases is that the latter do not require a primer to starting time making RNA. Once RNA polymerases are in the right place to start copying DNA, they just brainstorm making RNA by stringing together RNA nucleotides complementary to the DNA template.
This, of course, brings us to an obvious question- how do RNA polymerases "know" where to start copying on the Dna. Unlike the state of affairs in replication, where every nucleotide of the parental Dna must eventually be copied, transcription, equally we take already noted, simply copies selected genes into RNA at any given time.What indicates to an RNA polymerase where to offset copying Deoxyribonucleic acid to make a transcript? Signals in Deoxyribonucleic acid indicate to RNA polymerase where it should commencement (and end) transcription. These signals are special sequences in DNA that are recognized by the RNA polymerase or by proteins that help RNA polymerase decide where it should bind the DNA to first transcription. A Deoxyribonucleic acid sequence at which the RNA polymerase binds to beginning transcription is called a promoter.
A promoter is generally situated upstream of the gene that it controls. What this ways is that on the Deoxyribonucleic acid strand that the gene is on, the promoter sequence is "before" the gene. Remember that, by convention, DNA sequences are read from five' to 3'. So the promoter lies v' to the commencement betoken of transcription.
Also find that the promoter is said to "control" the gene it is associated with. This is because expression of the gene is dependent on the bounden of RNA polymerase to the promoter sequence to begin transcription. If the RNA polymerase and its helper proteins do not bind the promoter, the gene cannot be transcribed and it volition therefore, not exist expressed.

What is special nigh a promoter sequence? In an effort to answer this question, scientists looked at many genes and their surrounding sequences. It makes sense that because the aforementioned RNA polymerase has to bind to many unlike promoters, the promoters should have some similarities in their sequences. Certain enough, common sequence patterns were seen to be present in many promoters. We will first take a look at prokaryotic promoters. When prokaryotic genes were examined, the following features usually emerged:
- A transcription start site (this the base of operations in the DNA across from which the get-go RNA nucleotide is paired).
- A -x sequence: this is a half-dozen bp region centered about 10 bp upstream of the outset site. The consensus sequence at this position is TATAAT. In other words, if y'all count dorsum from the transcription starting time site, which past convention, is called the +ane, the sequence found at -10 in the majority of promoters studied is TATAAT).
- A -35 sequence: this is a sequence at about 35 basepairs upstream from the start of transcription. The consensus sequence at this position is TTGACA.
What is the significance of these sequences? It turns out that the sequences at -10 and -35 are recognized and bound by a subunit of prokaryotic RNA polymerase before transcription can begin.
The RNA polymerase of Eastward. coli, for case, has a subunit called the sigma (σ) subunit (or sigma gene) in improver to the core polymerase, which is the part of the enzyme that really makes RNA. Together, the sigma subunit and core polymerase make up what is termed the RNA polymerase holoenzyme. The sigma subunit of the polymerase can recognize and bind to the -ten and -35 sequences in the promoter, thus positioning the RNA polymerase at the right place to initiate transcription. In one case transcription begins, the core polymerase and the sigma subunit carve up, with the core polymerase continuing RNA synthesis and the sigma subunit wandering off to escort another core polymerase molecule to a promoter. The sigma subunit can be thought of as a sort of usher that leads the polymerase to its "seat" on the promoter.
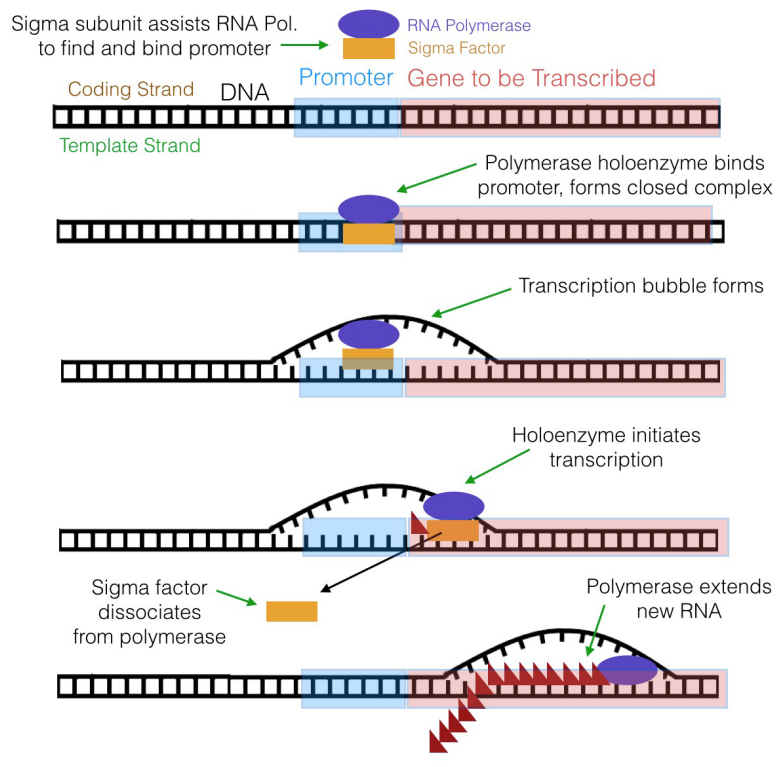
As already mentioned, an RNA chain, complementary to the Deoxyribonucleic acid template, is built by the RNA polymerase by the joining of the 5' phosphate of an incoming ribonucleotide to the 3'OH on the last nucleotide of the growing RNA strand. How does the polymerase know where to stop? A sequence of nucleotides called the terminator is the indicate to the RNA polymerase to stop transcription and dissociate from the template.
Although the process of RNA synthesis is the same in eukaryotes as in prokaryotes, in that location are some additional issues to keep in mind in eukaryotes. One is that in eukaryotes, the DNA template exists every bit chromatin, where the DNA is tightly associated with histones and other proteins. The "packaging" of the DNA must therefore exist opened up to allow the RNA polymerase admission to the template in the region to exist transcribed.
A second difference is that eukaryotes accept multiple RNA polymerases, not one as in bacterial cells. The different polymerases transcribe different genes. For instance, RNA polymerase I transcribes the ribosomal RNA genes, while RNA polymerase 3 copies tRNA genes. The RNA polymerase we will focus on most is RNA polymerase II, which transcribes protein-coding genes to make mRNAs.
All three eukaryotic RNA polymerases need boosted proteins to help them get transcription started. In prokaryotes, RNA polymerase past itself can initiate transcription (recollect that the sigma subunit is a subunit of the prokaryotic RNA polymerase). The additional proteins needed by eukaryotic RNA polymerases are referred to as transcription factors.
Finally, in eukaryotic cells, transcription is separated in infinite and time from translation. Transcription happens in the nucleus, and the mRNAs produced are candy further before they are sent into the cytoplasm. Protein synthesis (translation) happens in the cytoplasm. In prokaryotic cells, mRNAs can be translated as they are coming off the DNA template, and because there is no nucleus, transcription and poly peptide synthesis occur in a single cellular compartment.
Like genes in prokaryotes, eukaryotic genes also have promoters. Eukaryotic promoters usually accept a TATA box, a sequence about 25 base pairs upstream of the offset of transcription that is recognized and jump past proteins that help the RNA polymerase to position itself correctly to begin transcription. (Some eukaryotic promoters lack TATA boxes, and have, instead, other recognition sequences to help the RNA polymerase find the spot on the DNA where it spot on the DNA where it binds and initiates transcription.)
We noted earlier that eukaryotic RNA polymerases need additional proteins to bind promoters and first transcription. What are these additional proteins that are needed to start transcription? General transcription factors are proteins that assistance eukaryotic RNA polymerases find transcription start sites and initiate RNA synthesis. We will focus on the transcription factors that help RNA polymerase 2. These transcription factors are named TFIIA, TFIIB and and then on (TF= transcription factor, Two=RNA polymerase II, and the letters distinguish individual transcription factors).
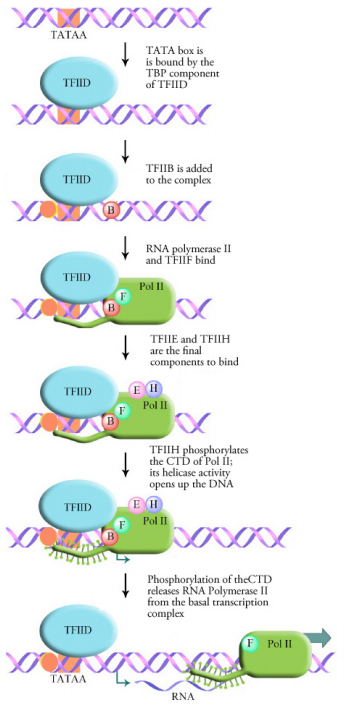
Transcription in eukaryotes requires the full general transcription factors and the RNA polymerase to form a complex at the TATA box called the basal transcription complex or transcription initiation complex. This is the minimum requirement for any gene to be transcribed. The showtime step in the formation of this complex is the binding of the TATA box by a transcription factor chosen the TATA Binding Poly peptide or TBP. Binding of the TBP causes the Deoxyribonucleic acid to bend at this spot and take on a structure that is suitable for the binding of additional transcription factors and RNA polymerase. As shown in the figure at left, a number of different general transcription factors, together with RNA polymerase (Pol II) form a complex at the TATA box.
The final step in the associates of the basal transcription circuitous is the binding of a full general transcription cistron called TFIIH. TFIIH is a multifunctional protein that has helicase activity (i.e., it is capable of opening upwardly a DNA double helix) as well as kinase activity. The kinase activity of TFIIH adds a phosphate onto the C-terminal domain (CTD) of the RNA polymerase. This phosphorylation appears to exist the bespeak that releases the RNA polymerase from the basal transcription complex and allows information technology to motility forwards and brainstorm transcription.
Either DNA strand can exist a template
The promoter is the sequence of Dna that encodes the information about where to begin transcription for each gene. Depending on the promoter, either strand of Dna can be used as the template strand.
Watch this video to see how either strand of DNA can be used equally a template for different genes on the same chromosome.
template vs. non-template strands summary
- The template strand is the 1 that RNA polymerase uses equally the ground to build the RNA. This strand is likewise called the non-coding strand or the antisense strand.
- The not-template strand has the identical sequence of the RNA (except for the substituion of U for T). This strand is as well chosen the coding strand or sense strand.
Major Types of Cellular RNA
Cells make several different kinds of RNA:
- mRNAs that code for proteins
- rRNAS that form role of ribosomes
- tRNAs that serve equally adaptors betwixt mRNA and amino acids during translation
- MicroRNAs that regulate gene expression
- Other pocket-sized RNAs that have a diversity of functions.
Contributors and Attributions
What Is The Promoter, Terminator, Template Strand, Non-template Strand,
Source: https://bio.libretexts.org/Courses/University_of_Arkansas_Little_Rock/Genetics_BIOL3300_%28Fall_2021%29/Genetics_Textbook/02:_Central_Dogma/2.01:_Overview_of_Transcription
Posted by: tomczaksayint.blogspot.com


0 Response to "What Is The Promoter, Terminator, Template Strand, Non-template Strand"
Post a Comment